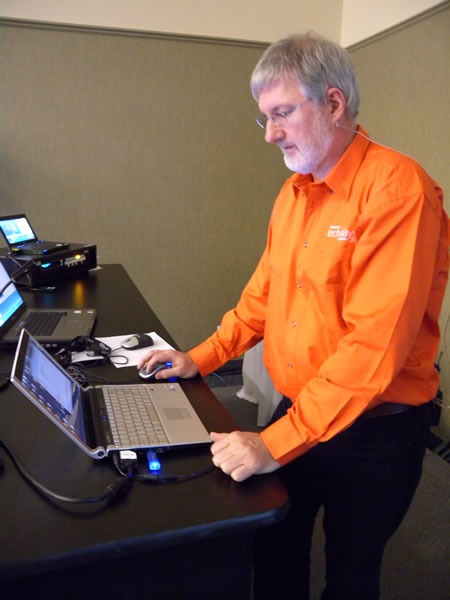
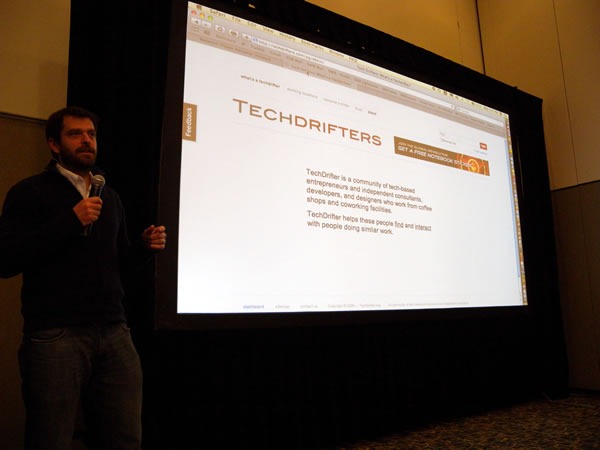
It’s hard to believe, but the seven-city cross-Canada tour known as TechDays 2009 is over. We had the last one – TechDays Winnipeg – on Tuesday and Wednesday of last week. Here are some photos I shot during the event.
The Day Before TechDays Winnipeg
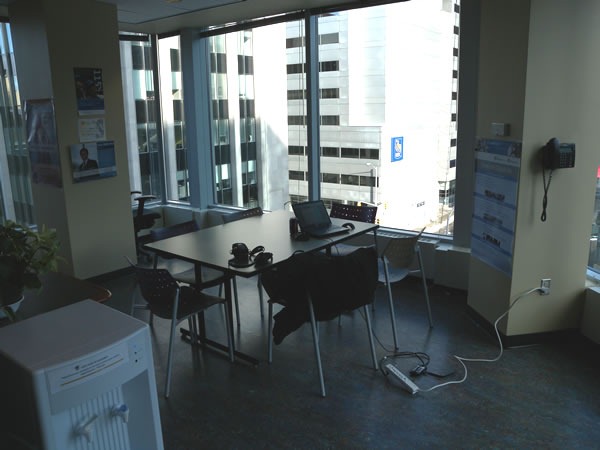
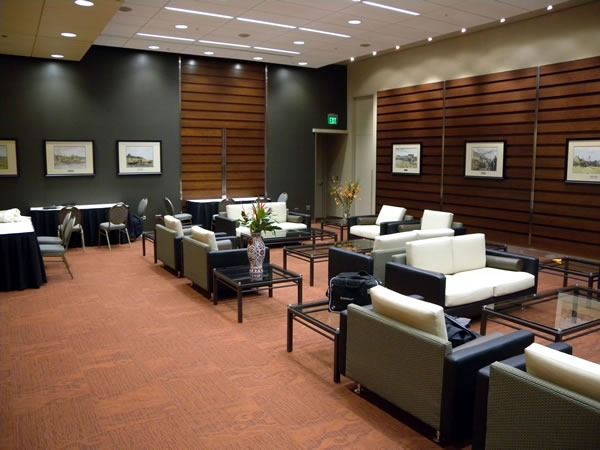
Of all the TechDays venues, I would have to hand the “swankiest speaker prep room” award to the Winnipeg Convention Centre, with its wood panelling, private washrooms, loads of closet space, plentiful tables, very comfortable leatherette couches and all-round 1980s styling. I can imagine a young Flock of Seagulls or Duran Duran hanging out here after a show, entertaining groupies:

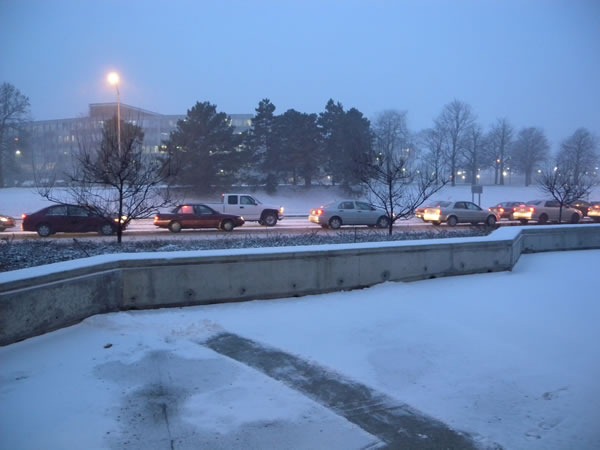
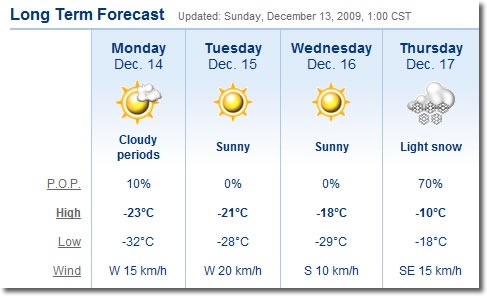
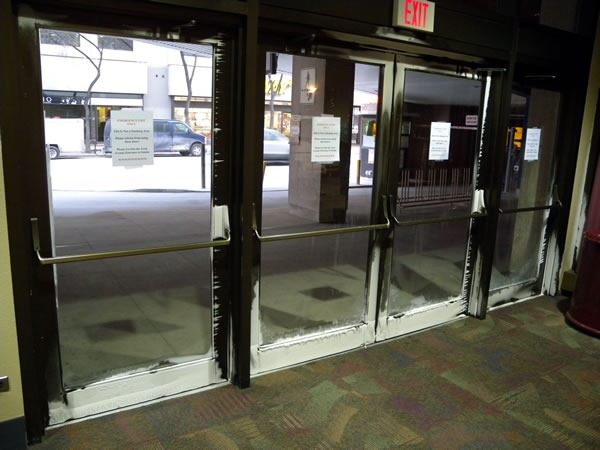
During the holidays, many people like to decorate their storefront and home windows with fake spray-on frost. In Winnipeg, where the temperatures were hovering around –35 degrees C (-31 degrees F), you don’t need that stuff – they’ve got the real thing! Here are the side doors on the ground floor of the Winnipeg Convention Centre:

Here’s a closer look:
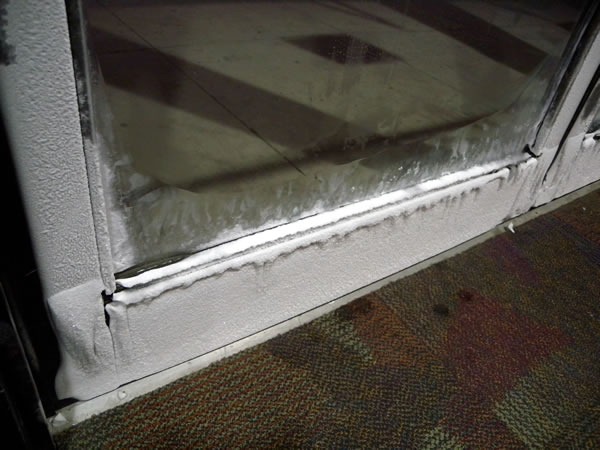

And just for kicks, an even closer one.

I must tip my hat to the people of Winnipeg for toughing out those kinds of temperatures, year after year.
The Convention Centre had a secret stash of Christmas trees, ready to be deployed at a moment’s notice:

One of the perks of being a TechDays Track Lead is that nobody asks questions when you rearrange the signs for an art shot:

Day 1
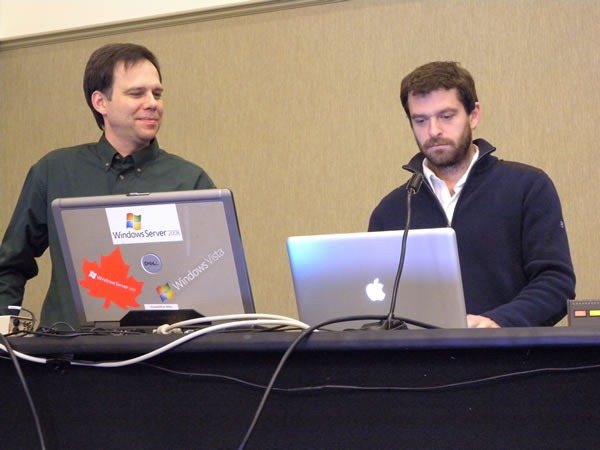
John Bristowe, track lead for the green-shirted Developer Fundamentals and Best Practices track, just had another baby, so he was tied up with Dad duties (congrats, John and Fiona!). I donned a green shirt took over as acting track lead for his track and recruited D’Arcy Lussier to host my track, the orange-shirted Developing for the Microsoft-Based Platform track.
The first speaker for Developer Fundamentals and Best Practices was Jeremy Wiebe, who presented the very popular Tips and Tricks for Visual Studio session:

How popular was it? Popular enough that people were overflowing out of the rows:
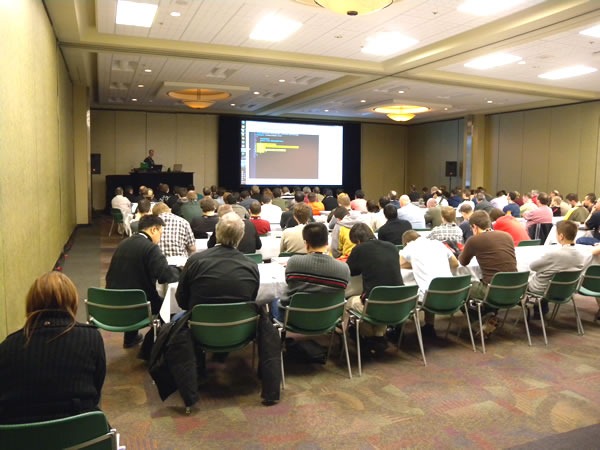

…and we even had to drag in some extra chairs to create a new row at the back:
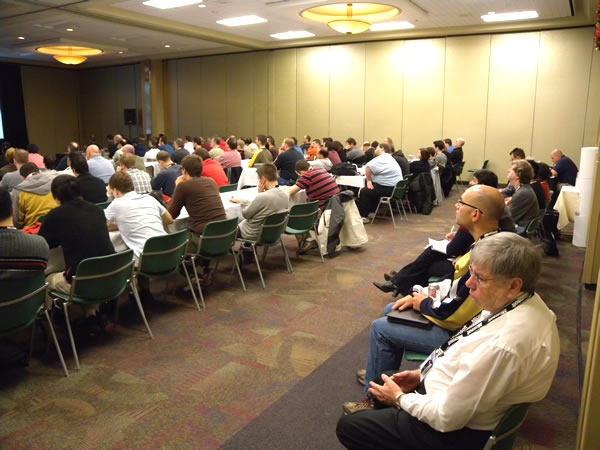

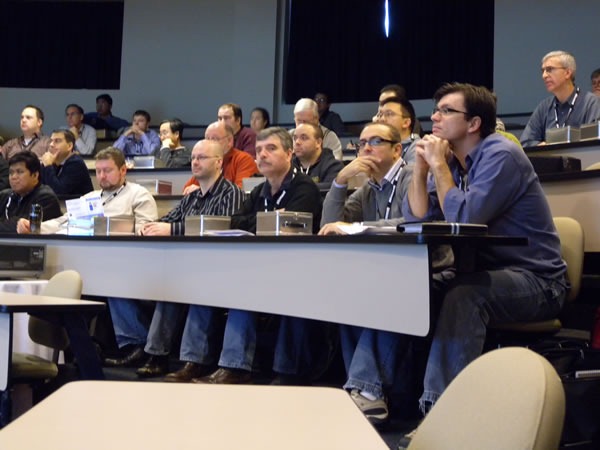
This was an attentive crowd. There were a lot of “I didn’t know you could do that in Visual Studio!”-type reactions.
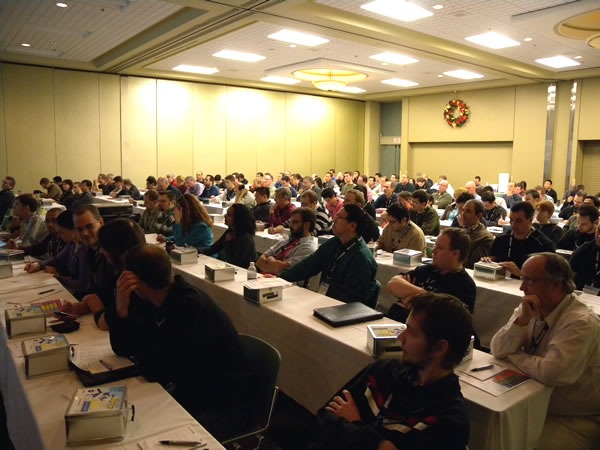

The second session of the day was given by Dylan Smith: Test-Driven Development Techniques:
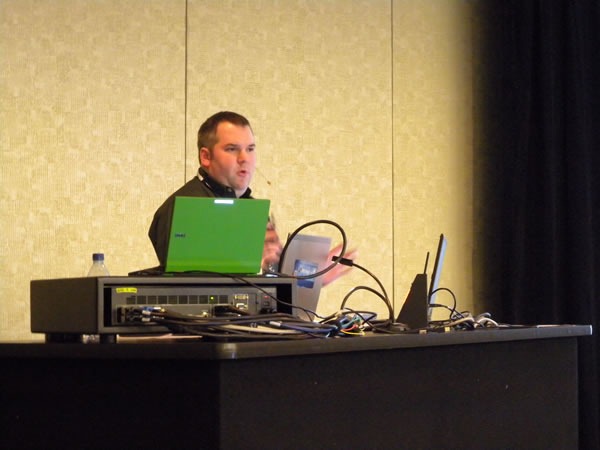

Once again, a good crowd.
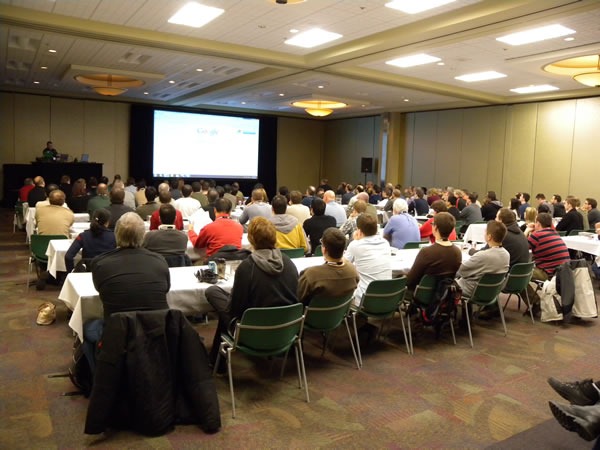

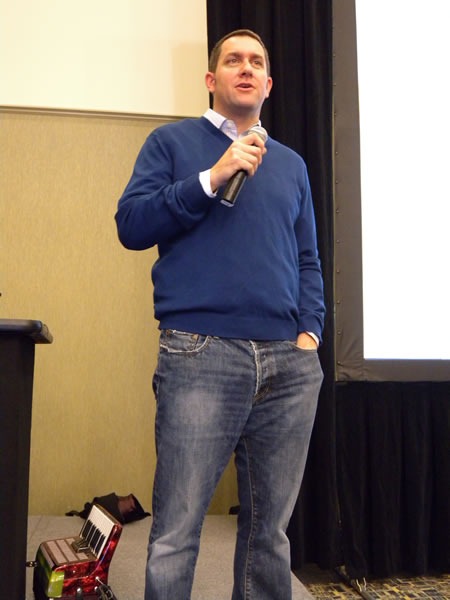
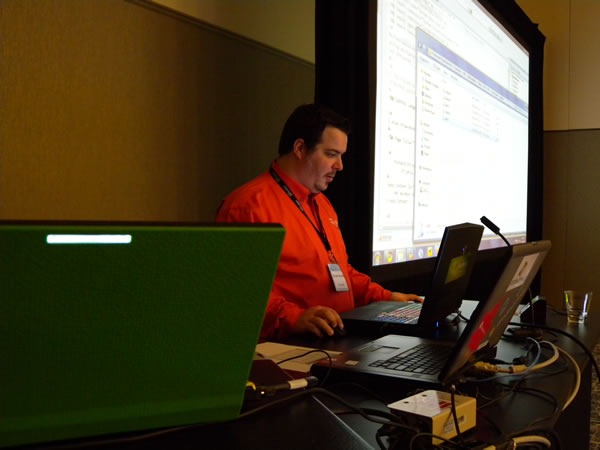
During lunch, my coworker, IT pro evangelist Rick Claus and I did a presentation on some of the new features in Office 2010, with me showing off some of the new graphics goodness in PowerPoint 2010:
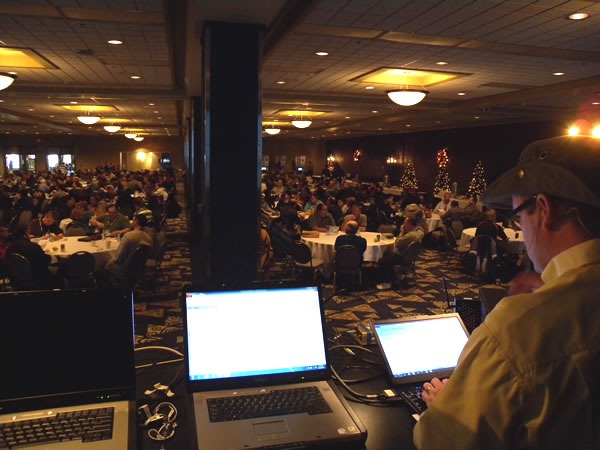

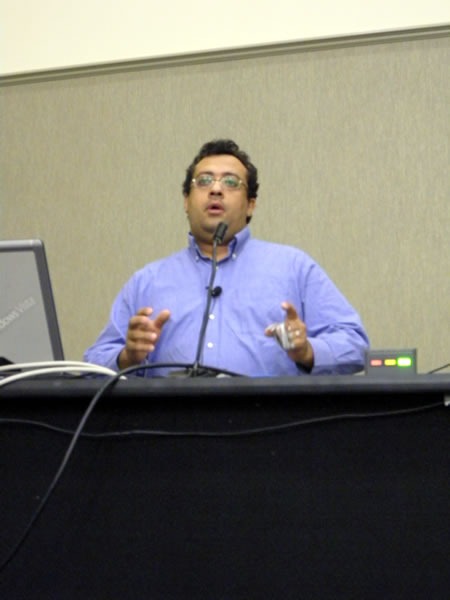
The sessions resumed in the afternoon with Uwe Schmitz talking about Patterns for the Rest of Us. I was a bit surprised at how few hands went up when I asked how many people had read or even attempted to read Design Patterns by the “Gang of Four”.

Most of Uwe’s audience was in the same room as he was:
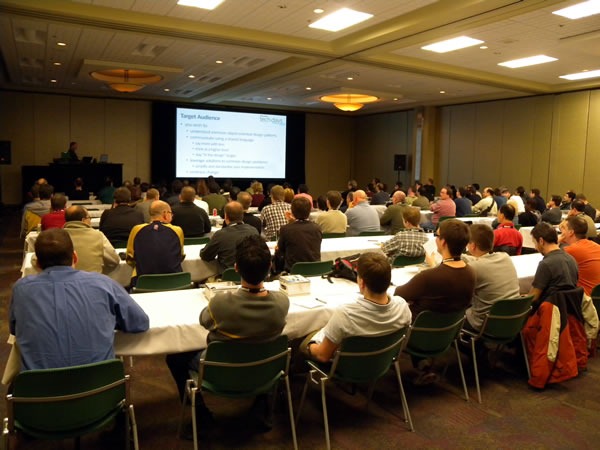

But one guy, whose back was made sore by the conference room chairs, took a clever approach. We broadcast all sessions’ projections and audio on a monitor outside every room, so he took one of the comfy chairs in the hallway outside and set himself up for some living room-style viewing:
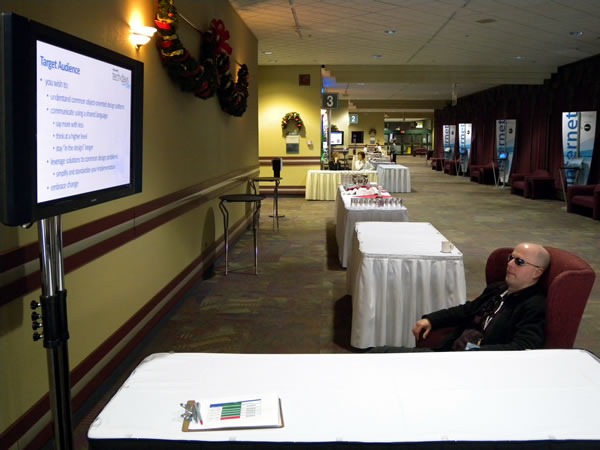

I told him that with his sunglasses and the way he was seated, he reminded me of the old ads for Maxell tapes from the 1980s:

After Uwe was Dave Harris, who presented A Strategic Comparison of Data Access Technologies from Microsoft:

Day 2
The outside temperature improved for the second day: it became a relatively balmy –20 degrees C (-4 degrees F). What a difference 15 degrees makes!
The first session was Practical Web Testing and was delivered by the team of Tyler Doerksen and Robert Regnier:

I stepped out to drop in on the track which I had put together, my orange-shirted Developing for the Microsoft-Based Platform track. While the Developer Fundamentals and Best Practices track typically had big draws on Day 1, Day 2 is when the Platform track brought in the crowds:
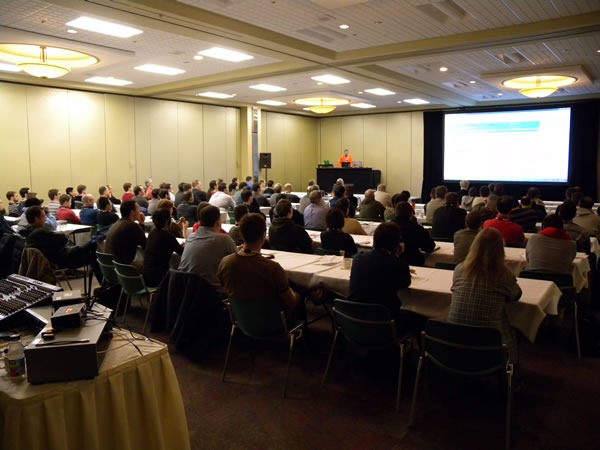

The session was the popular Introducing ASP.NET MVC, and in Winnipeg, it was delivered by Kelly Cassidy:
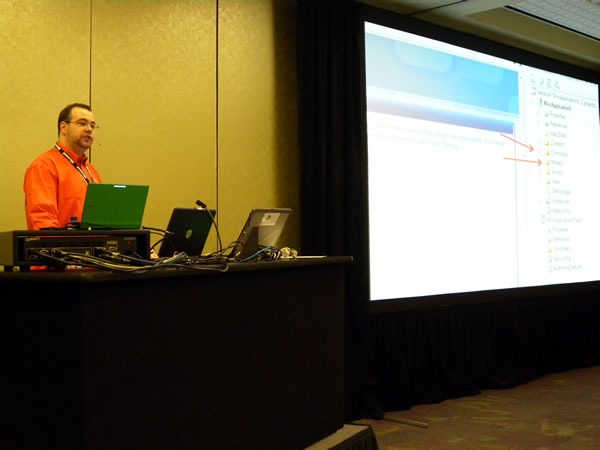

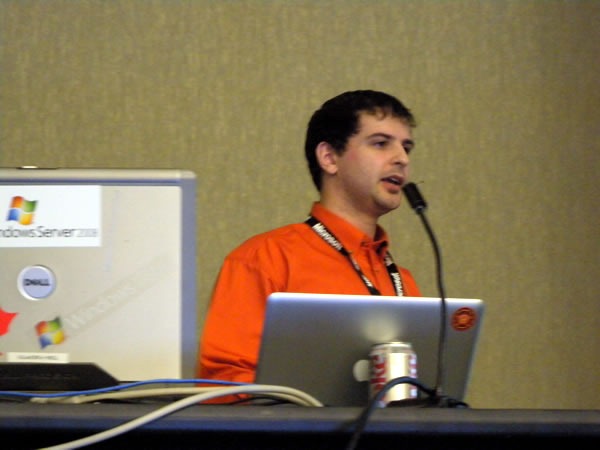
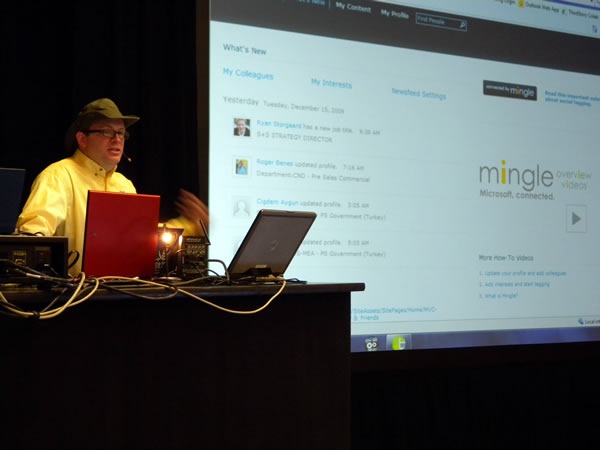
An unfortunate set of circumstances speaker shortages and cancellations means that Rick had to deliver all the presentations for day 2 of his track, Servers, Security and Management. That’s 300 minutes in total behind the lectern. It’s quite fortunate that he knows his stuff and that his theatre training makes him an excellent presenter:
Meanwhile, back at the green track, Aaron Kowall presented Better Software Change and Configuration Management Using TFS:
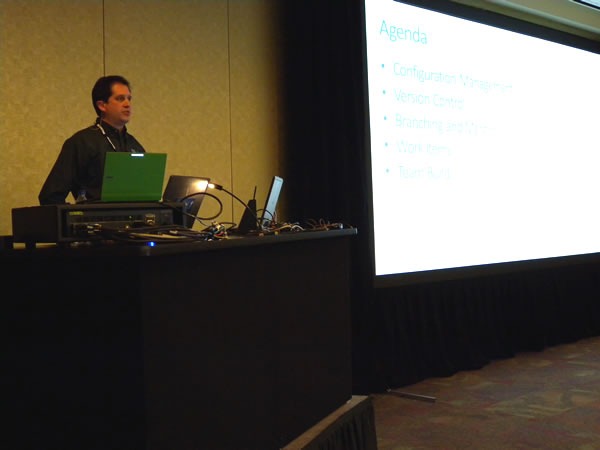

During his session, he presented a very important truth: Build automation is not just merely pressing “F5”:

At lunch, Rick hosted a demo showdown between me (representing developers) and my coworker, IT Pro Evangelist Rodney Buike, trying to determine who could do the better Windows 7 demos. I won, thanks in part to my demo of the most obscure Windows features: the Private Character Editor.
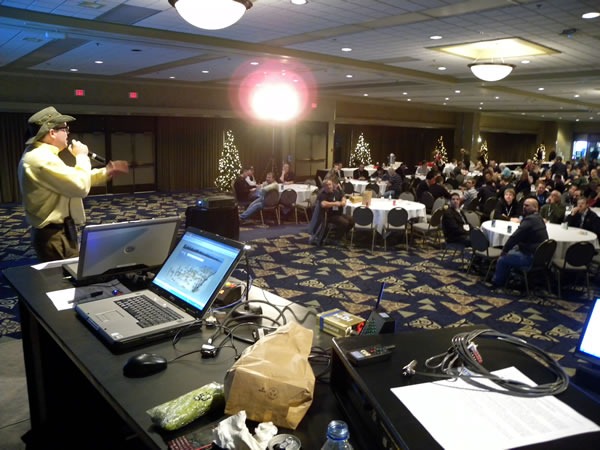

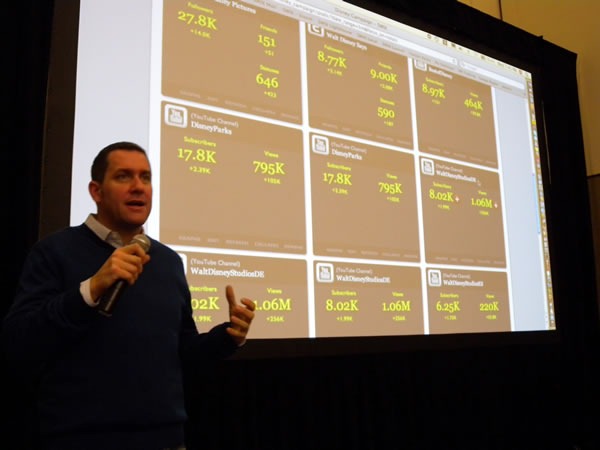
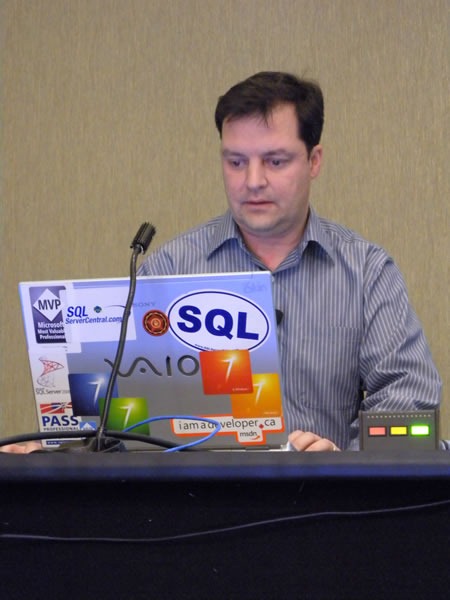
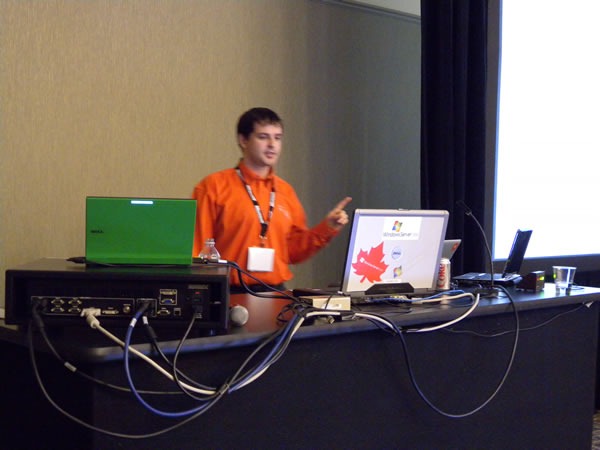
Joel Semeniuk needs no introduction. I simply told the audience that “Joel has forgotten more about Team System than I will ever learn. Besides, what I know about Team System can be summarized in the two words ‘jack’ and ‘poop’.” Here’s Joel in action, presenting Metrics That Matter: Using Team System for Process Improvement:

I love this shot of Joel – he looks like a general addressing his own private banana republic:
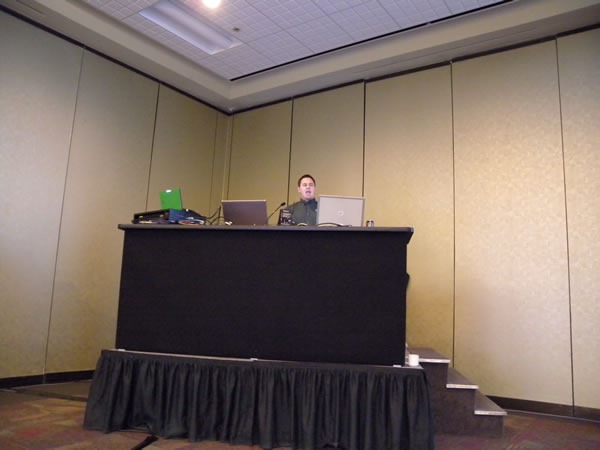

A closer look:
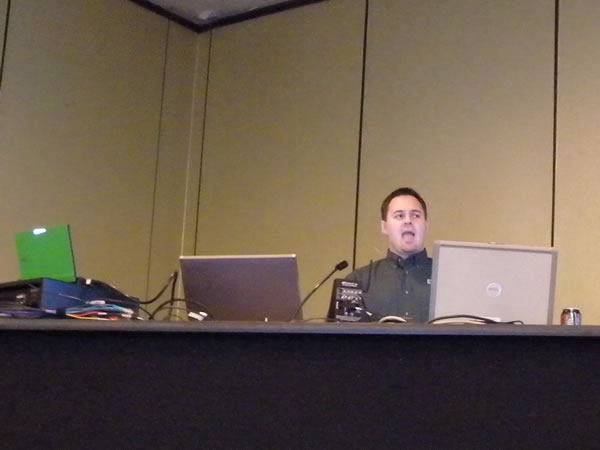

The most popular afternoon snack was served between the third and fourth sessions of Day 2: Canada’s favourite snack – donuts!
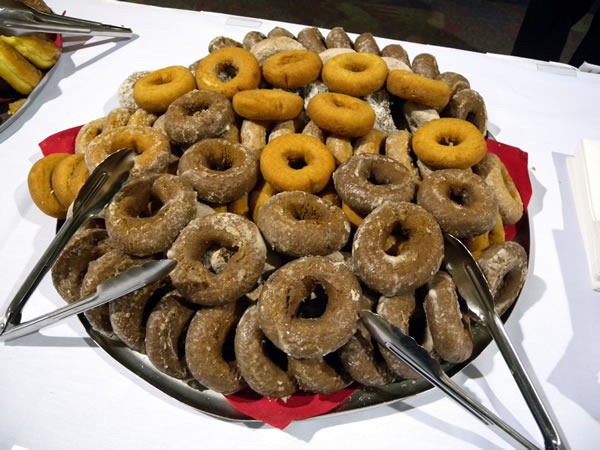

My SD card corrupted the photos of the last speaker of the day, Steve Porter, who did a fine job presenting his session, Database Change Management with Team System. My apologies, Steve!
And finally, to make up for the fact that I did not properly capture D’Arcy Lussier’s hair — an asset in which he takes great pride — in yesterday’s video interview, I now present a close-up shot of his coiff:
My thanks to everyone at TechDays Winnipeg – attendees, speakers, staff and organizers – for making it an great way to close out the tour!
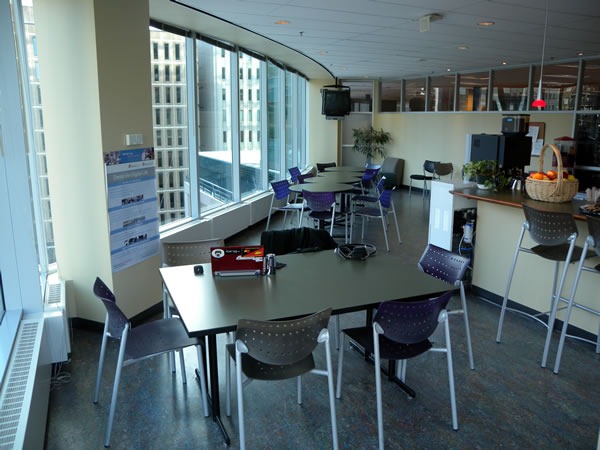
 Microsoft Ottawa’s Kitchen. It has a decent view.
Microsoft Ottawa’s Kitchen. It has a decent view.